The Plant Cell 21: 3803-3822 (2009)
Ethylene interacts with abscisic acid to regulate endosperm rupture during germination: a comparative approach using Lepidium sativum and Arabidopsis thaliana [W][OA]
Warwick Horticulture Research International (HRI), Warwick University, Wellesbourne, Warwick CV35 9EF, United Kingdom (Ka.M., C.S.C.C., J.R.L., W.E.F.-S.)
Palacky University and Institute of Experimental Botany Academy of Sciences of the Czech Republic, Laboratory of Growth Regulators, CZ-78371 Olomouc, Czech Republic (V.T., M.S.)
Université Pierre et Marie Curie-Paris 6, Germination et Dormance des Semences, UR5, Site d'Ivry, F-75005 Paris, France (F.C.)
Received July 23, 2009; Returned for revision October 12, 2009; Accepted November 17, 2009; Published December 18, 2009
www.plantcell.org/cgi/doi/10.1105/tpc.109.070201
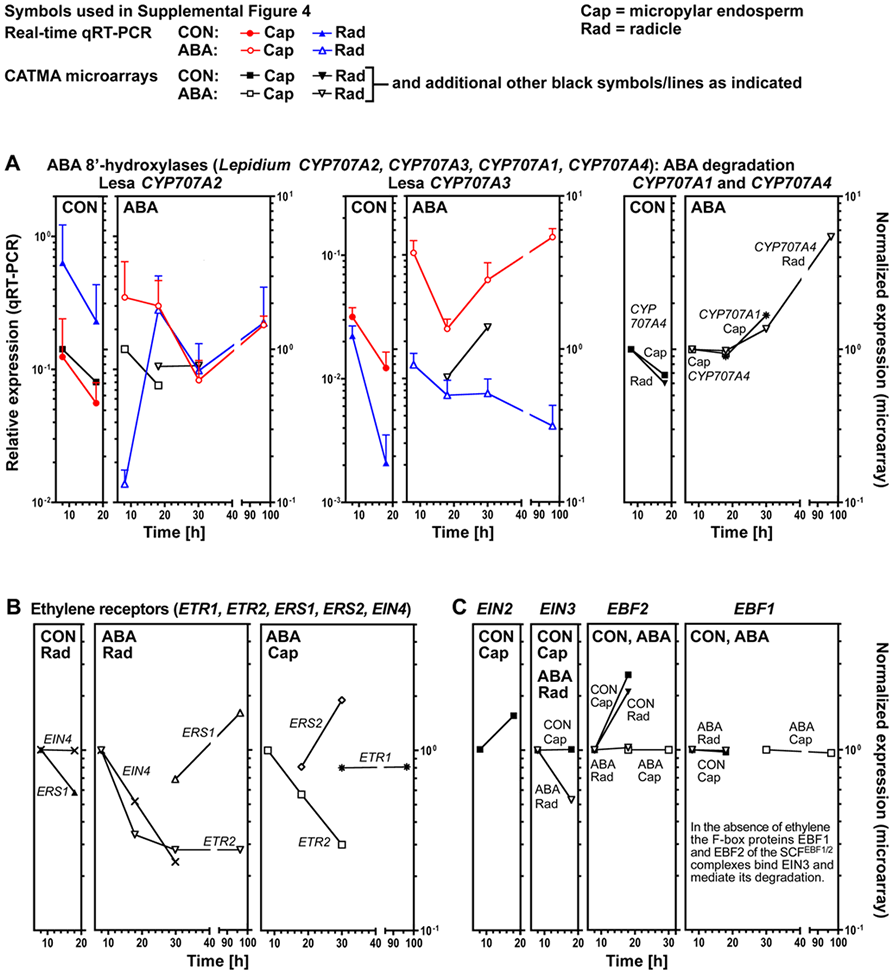
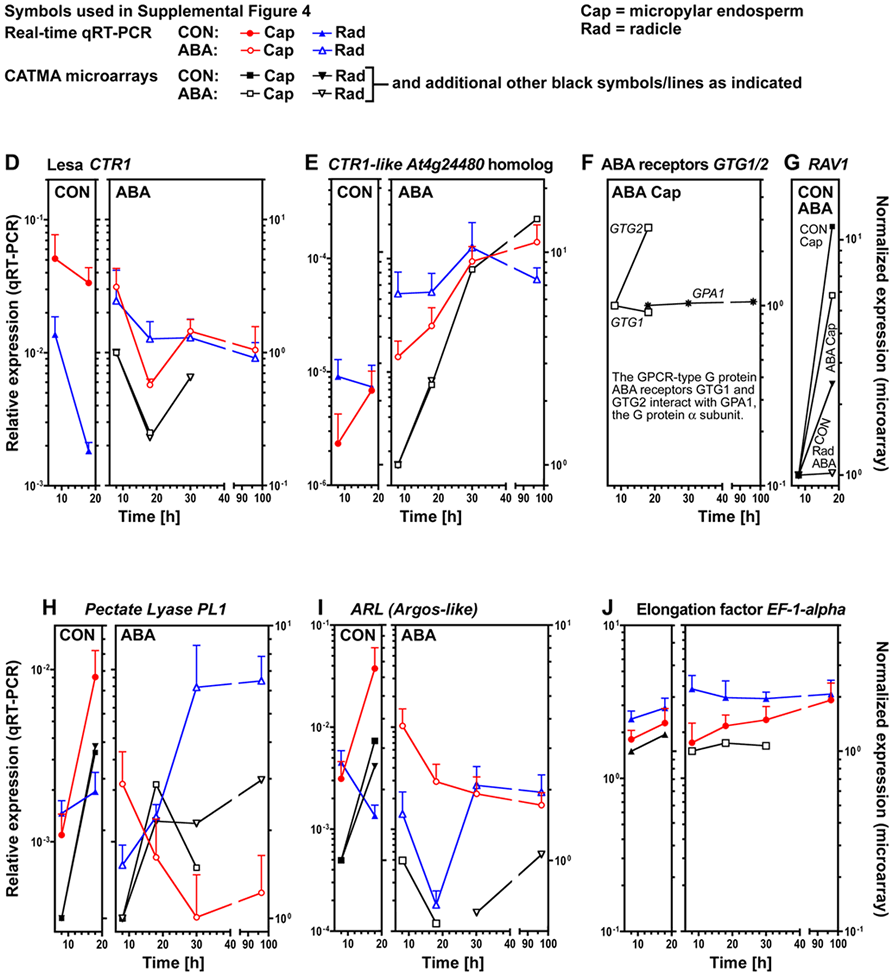
Supplemental Figure 4. Analysis of transcript expression by qRT-PCR and microarray analysis in specific seed tissues of Lepidium sativum FR1 during germination.
Time course transcript expression data are presented for endosperm caps (Cap) and radicles (Rad) dissected from seeds incubated at 24ºC in continuous light in medium without (CON) or with 10 µM ABA added. For qRT-PCR relative deltadeltaCt expression values based on the comparison with validated constitutive transcripts are presented. Lepidium transcripts named with the prefix 'Lesa' were analysed by qRT-PCR primers designed on the basis of their cloned cDNA sequences. Lepidium transcripts without this prefix in their name were analysed with a qRT-PCR primer design based on Arabidopsis cDNA sequences. Primer sequences for the qRT-PCR are presented in Supplemental Table 1 online.
(A) ABA 8'-hydroxylases: Four CYP707A genes are known in Arabidopsis and all provided expression results in the Lepidium seed arrays. Two Lepidium cDNAs were cloned (Lepidium CYP707A2 and CYP707A3) and on the basis of their sequence analyses, represent putative Lepidium orthologs of Arabidopsis CYP707A2 and CYP707A3, respectively.
(B) Ethylene receptors.
(C) Ethylene signaling components.
(D) Ethylene signaling repressor CTR1 (Constitutive Triple Response1); the cDNA of the putative Lepidium ortholog CTR1 was cloned.
(E) CTR1-like serine/threonine protein kinase.
(F) GPCR-type G protein ABA receptors GTG1 and GTG2 and their interacting G protein a subunit GPA1.
(G) RAV1, an AP2/EREBP-type transcription factor with an ABI3/VP1-like domain.
(H) PL1, Pectate lyase1. (I) ARL, Argos-like, putative cell expansion gene.
(J) EF-1-alpha, Transcription elongation factor 1-alpha.
Mean values +SE of four independent biological RNA samples obtained from 1000 endosperm caps or 100 radicles from seeds with ruptured testa, but intact endosperm are presented for the qRT-PCR results. Normalized microarray differences are presented for comparison.
Synopsis: Tissue weakening of the endosperm is required to allow radicle protrusion during seed germination. Cross-species work including tissue-specific transcriptome analysis and biomechanical measurement of endosperm weakening provided a new mechanistic model that explains how ethylene promotes seed germination and counteracts the inhibition of endosperm cap weakening by abscisic acid.
| Article in PDF format (1.8 MB) Supplemental data file (1.8 MB) Abstract of Plant Cell 2009 |
|
|
|
The Seed Biology Place |
Webdesign Gerhard Leubner 2000 |